Self-organising principles of tissue formation
The human body consists of around 30 trillion cells. These cells self-organise into different types of tissues, all of which display striking architectures, including compartmentalisation into proliferative zones and regions with specialised cells. Tissue formation is regulated by a plethora of factors, including cellular shape and polarisation, cell-cell interactions, signalling events and microenvironments.
Frequently, tissue self-organisation is derailed during regeneration or cancer development, and the aim of many therapies is to restore normal tissue homeostasis by reversing cellular dysfunction or eliminating diseased cells. Understanding the principles of self-organisation in tissues is thus crucial for many areas of biomedicine.
To understand and control tissue formation, self-renewal and function, we need to know, for example, how cell division and its orientation affect cell fate, differentiation trajectories and tissue morphogenesis. We need to get insight into how specific alterations in cell architecture, such as cytoskeletal re-organisation or organelle positioning and dynamics contribute to cell differentiation, how robust these processes are and whether and how they can be steered. We also need to understand the interplay between cellular and subcellular events on different scales: for example, whether organelle positioning during or after cell division impacts cell fate.
Microscopy-based cell biological analysis of living tissues is the only way to address such questions.
Tissue perturbation, pathological transformation and repair
Tissues have a strong ability to regenerate when perturbed by mechanical, chemical, infectious or other assaults. For example, damaged epithelia can rapidly mount autonomous regenerative responses, resulting in cell de-differentiation, increased proliferative activity and cell migration, thus allowing restoration of tissue integrity. However, when deregulated, regenerative signals can also trigger disease, such as cancer and inflammation. So far, tissue alterations during repair as well as cancer initiation and progression have been mostly documented at the histological and transcriptional level.
Understanding tissue development by combining cell biological studies of organoids and tissues in vivo
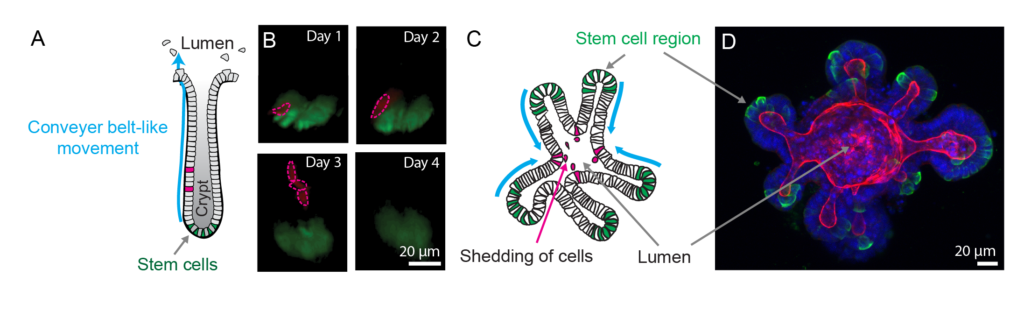
Organoids provide in vivo-like tissue-mimetic systems to explore central cell biology aspects of self-organising 3D tissues. Organoids are derived from adult mouse or human stem cells and mimic the stem cell niche [1], which self-organises into organ-like 3D structures that resemble the overall architecture and cellular composition/positioning of the tissue of origin. Organoids have been derived from many epithelia, including intestine, breast and airway, including samples from patients, and have been successfully applied to identify mechanisms of tissue formation and disease [2-4]. Importantly, organoids offer a unique opportunity for high-resolution imaging at different scales (tissue to cell to molecule). Their amenability to live cell microscopy, genetic and pharmacological manipulation renders them excellently suited for uncovering the integration of molecular and cellular function and malfunction in tissue development and disease. However, since organoids are cultured tissues that lack the complex environmental factors present within an organism, they need to be carefully validated by detailed comparisons to tissues in vivo.
References:
- Sato T et al., 2009. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature, 459: 262-5.
- Florian S et al., 2019. A human organoid system that self-organizes to recapitulate growth and differentiation of a benign mammary tumor. Proc Natl Acad Sci U S A, 116: 11444-11453.
- Jamieson PR et al., 2017. Derivation of a robust mouse mammary organoid system for studying tissue dynamics. Development, 144: 1065-1071. 11.
- Simian M et al., 2001. The interplay of matrix metalloproteinases, morphogens and growth factors is necessary for branching of mammary epithelial cells.Development, 128: 3117-31.
